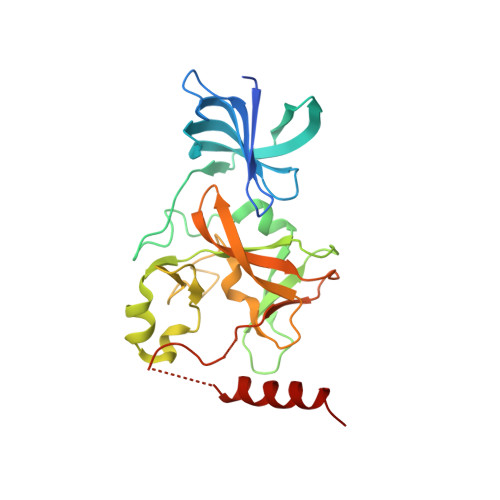
Identification of Cyproheptadine as an Inhibitor of SET Domain Containing Lysine Methyltransferase 7/9 (Set7/9) That Regulates Estrogen-Dependent Transcription
Takemoto, Y., Ito, A., Niwa, H., Okamura, M., Fujiwara, T., Hirano, T., Handa, N., Umehara, T., Sonoda, T., Ogawa, K., Tariq, M., Nishino, N., Dan, S., Kagechika, H., Yamori, T., Yokoyama, S., Yoshida, M.(2016) J Med Chem 59: 3650-3660
- PubMed: 27088648
- DOI: https://doi.org/10.1021/acs.jmedchem.5b01732
- Primary Citation of Related Structures:
5AYF - PubMed Abstract:
SET domain containing lysine methyltransferase 7/9 (Set7/9), a histone lysine methyltransferase (HMT), also methylates non-histone proteins including estrogen receptor (ER) α. ERα methylation by Set7/9 stabilizes ERα and activates its transcriptional activities, which are involved in the carcinogenesis of breast cancer. We identified cyproheptadine, a clinically approved antiallergy drug, as a Set7/9 inhibitor in a high-throughput screen using a fluorogenic substrate-based HMT assay. Kinetic and X-ray crystallographic analyses revealed that cyproheptadine binds in the substrate-binding pocket of Set7/9 and inhibits its enzymatic activity by competing with the methyl group acceptor. Treatment of human breast cancer cells (MCF7 cells) with cyproheptadine decreased the expression and transcriptional activity of ERα, thereby inhibiting estrogen-dependent cell growth. Our findings suggest that cyproheptadine can be repurposed for breast cancer treatment or used as a starting point for the discovery of an anti-hormone breast cancer drug through lead optimization.
Organizational Affiliation:
Chemical Genetics Laboratory, RIKEN , 2-1 Hirosawa, Wako, Saitama 351-0198, Japan.